Primer BLAST performs only a specificity check when a target template and both primers are provided. In the Primer Pair Specificity Checking Parameters section, select the appropriate source Organism and the smallest Database that is likely to contain the target sequence. These settings give the most precise results. Colony PCR is a convenient high-throughput method for determining the presence or absence of insert DNA in plasmid constructs. Individual transformants can either be lysed in water with a short heating step or added directly to the PCR reaction and lysed during the initial heating step.
Hello, I have this problem in checking the positive insert after cloning. I've already prepared the miniprep and tried digesting the plasmid with the suitable restriction enzymes. Somehow i cannot confirmed the diagnostics / interest fragments. Now, I'm just wondering is it possible for me to check the positive insert using the plasmid DNA (without using restriction enzymes) but by PCR (something like colony PCR)? Is this feasible? If so, can someone be kind enough to teach me how to do it?
What primer should i used?ThanksDesperately need helps: spices. Spices-before going down the PCR route, why was the restriction digest a failure/inconclusive? A restriction digest is far better way of confirming that your plasmid is properly built.
PCR simply says that the segment which your primers amply is correct. It does not say anything about rest of the plasmid. Often PCR is used first as a screen and later confirmed by restriction digest, and perhaps even sequencing.As for the PCR; you should try using one primer that binds to the vector and one primer that binds to the insert.
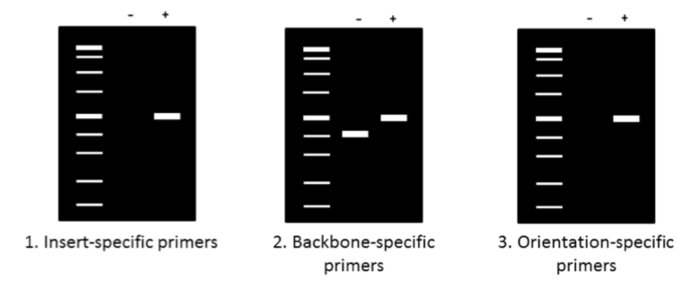
A PCR product will tell you insert in in your plasmid (at least the end which was sequenced). You may then want to check the other end of the fragment. Timjim-Thanks for all the advice. The reason i can't decide on the positive insert using restriction enzymes is that somehow for this particular experiment ( i don't know why. I never encounter this before), my heavier band/ fragment was very pale and my diagnostic fragment happened to be a little bit smaller (i would say not that small.
About 700ish bp & 1000is bp), but since the heavier band was already very blur. Searching for smaller fragments were almost impossible.
I tried to re-stain, destained my gels and yet it is still not helpful. So, that's the reason why i'm thinking to do PCR on my plasmid, at least just to give some idea whether i have insert or not. By the way, does anyone know what can cause this blurry fragment?Thanks againBlurry spices. Spices-a picture would help greatly.A blurry band could be1: DNA is degraded2: Too much DNA in well3: Too much salt in loading sample (often from digestion buffer)4: Gel was run too fast5: Gel has too much salt (problems in preparing the gel)6: Gel is old and contaminated.
Resulting in the DNA being degraded whiles it is running7: Loading buffer is contaminated, resulting in DNA being degraded whiles running.8: For small fragments (below 1000bp), gel was run too slowly, allowing DNA to diffuse.
